Research at the Baca Lab
Examples of recent project include:
Studying cancer epigenomes using liquid biopsy (Nature Medicine)
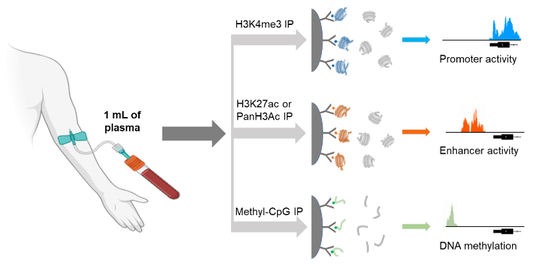
Circulating tumor DNA (ctDNA) assays are increasingly used to inform clinical decisions in cancer care, but they have limited ability to identify the transcriptional programs that govern cancer phenotypes and their dynamic changes during the course of disease. To address these limitations, we collaborated with the lab of Dr. Matthew Freedman to developed a method for comprehensive epigenomic profiling of cancer from 1 mL of patient plasma. Using an immunoprecipitation-based approach targeting histone modifications and DNA methylation, we measured 1,268 epigenomic profiles in plasma from 433 individuals with one of 15 cancers. Our assay provides a robust proxy for transcriptional activity, allowing us to infer the expression levels of diagnostic markers and drug targets, measure the activity of therapeutically targetable transcription factors and detect epigenetic mechanisms of resistance. Work is ongoing to test the clinical utility of this approach and determine how epigenetic changes in cancer drive treatment resistance.
Studying cancer epigenomes using liquid biopsy (Nature Medicine)
Circulating tumor DNA (ctDNA) assays are increasingly used to inform clinical decisions in cancer care, but they have limited ability to identify the transcriptional programs that govern cancer phenotypes and their dynamic changes during the course of disease. To address these limitations, we collaborated with the lab of Dr. Matthew Freedman to developed a method for comprehensive epigenomic profiling of cancer from 1 mL of patient plasma. Using an immunoprecipitation-based approach targeting histone modifications and DNA methylation, we measured 1,268 epigenomic profiles in plasma from 433 individuals with one of 15 cancers. Our assay provides a robust proxy for transcriptional activity, allowing us to infer the expression levels of diagnostic markers and drug targets, measure the activity of therapeutically targetable transcription factors and detect epigenetic mechanisms of resistance. Work is ongoing to test the clinical utility of this approach and determine how epigenetic changes in cancer drive treatment resistance.
Reprogramming of FOXA1 DNA binding in treatment-emergent neuroendocrine prostate cancer (Nature Communications)
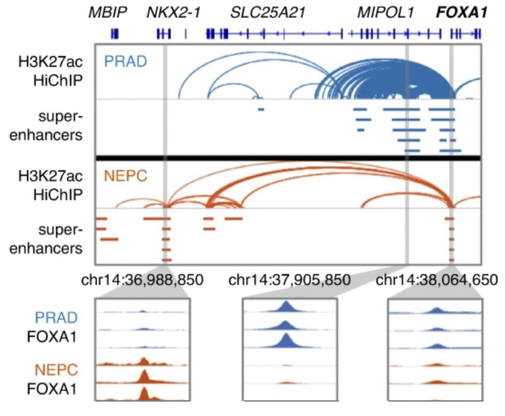
Despite the rapidly growing number of effective cancer treatments, most advanced cancers stop responding to therapy eventually. A poorly understood mechanism of treatment resistance in prostate cancer is lineage plasticity. Lineage plasticity occurs when prostate adenocarcinoma (PRAD) cells transform into neuroendocrine prostate cancer (NEPC). NEPC is an aggressive cancer that does not respond to inhibition of androgen signaling - the mainstay of prostate cancer treatment. We investigated the epigenomic basis of this resistance mechanism by profiling histone modifications in NEPC and PRAD using ChIP-seq. We identified a vast network of regulatory elements that are activated in NEPC by the transcription factor FOXA1, a cofactor of the Androgen Receptor (AR). Despite shedding its dependence on AR signaling, NEPC requires FOXA1 for survival. Expression of NEPC lineage transcription factors in PRAD re-directs FOXA1 binding to NEPC-specific regulatory elements, where it drives neuroendocrine gene expression programs. This study found that FOXA1 is essential for the survival of NEPC. More broadly, this work provides a principled approach to identifying cancer dependencies by profiling cancer epigenomes.
Despite the rapidly growing number of effective cancer treatments, most advanced cancers stop responding to therapy eventually. A poorly understood mechanism of treatment resistance in prostate cancer is lineage plasticity. Lineage plasticity occurs when prostate adenocarcinoma (PRAD) cells transform into neuroendocrine prostate cancer (NEPC). NEPC is an aggressive cancer that does not respond to inhibition of androgen signaling - the mainstay of prostate cancer treatment. We investigated the epigenomic basis of this resistance mechanism by profiling histone modifications in NEPC and PRAD using ChIP-seq. We identified a vast network of regulatory elements that are activated in NEPC by the transcription factor FOXA1, a cofactor of the Androgen Receptor (AR). Despite shedding its dependence on AR signaling, NEPC requires FOXA1 for survival. Expression of NEPC lineage transcription factors in PRAD re-directs FOXA1 binding to NEPC-specific regulatory elements, where it drives neuroendocrine gene expression programs. This study found that FOXA1 is essential for the survival of NEPC. More broadly, this work provides a principled approach to identifying cancer dependencies by profiling cancer epigenomes.
A blood test for neuroendocrine transformation of prostate cancer based on plasma DNA methylation (Clinical Cancer Research)
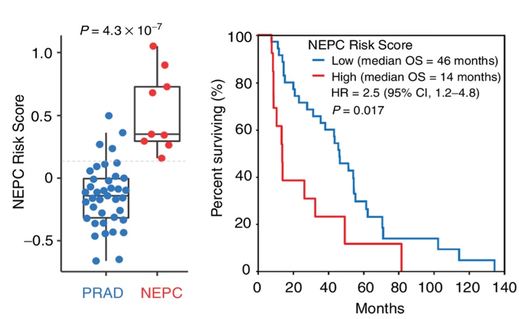
Detecting transformation of prostate adenocarcinoma (PRAD) into neuroendocrine prostate cancer (NEPC) has important prognostic and therapeutic implications. A blood biopsy-based approach to NEPC diagnosis would be clinically useful, allowing patients to receive appropriate treatment for this aggressive disease. We developed an assay based on circulating cell-free DNA methylation to detect the transformation of PRAD into NEPC from a blood draw. Applying the assay to plasma from two independent cohorts of men with mCRPC accurately identified men with NEPC. In both cohorts, high NEPC Risk Scores were associated with worse overall survival. This assay will fill a critical unmet need for non-invasive NEPC diagnostics.
Detecting transformation of prostate adenocarcinoma (PRAD) into neuroendocrine prostate cancer (NEPC) has important prognostic and therapeutic implications. A blood biopsy-based approach to NEPC diagnosis would be clinically useful, allowing patients to receive appropriate treatment for this aggressive disease. We developed an assay based on circulating cell-free DNA methylation to detect the transformation of PRAD into NEPC from a blood draw. Applying the assay to plasma from two independent cohorts of men with mCRPC accurately identified men with NEPC. In both cohorts, high NEPC Risk Scores were associated with worse overall survival. This assay will fill a critical unmet need for non-invasive NEPC diagnostics.
An epigenomic framework for understanding inherited risk of cancer (Nature Genetics)
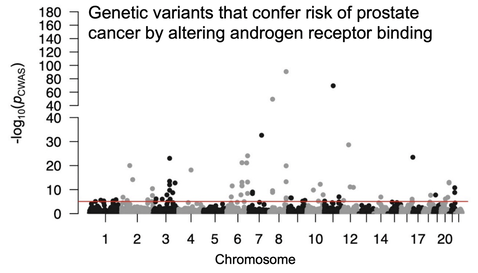
A substantial portion of people's risk of developing cancer is due to genetics. Much of the genetic variation underlying cancer risk affects regulatory elements (e.g., enhancers and promoters) that control gene expression. To understand how this variation in regulatory DNA confers risk of prostate cancer, we developed the cistrome-wide association study (CWAS). CWAS is a framework for identifying variants that impact traits and disease through their effects on chromatin state. CWAS associates the genetic determinants of cistromes (the genome-wide profiles of transcription factor binding sites or histone modifications) with phenotypes using summary statistics from genome-wide association studies (GWAS). We performed a CWAS of prostate cancer using a reference panel of 307 prostate cistromes from 165 individuals. CWAS identified hundreds of prostate cancer risk loci where regulatory element activity or transcription factor binding explain most of the inherited risk identified by GWAS. In addition, CWAS boosts discovery power in modestly sized GWAS, allowing us to identify novel genetic associations mediated through AR binding. This allowed us to discover a SNP associated with resistance to prostate cancer therapy from a clinical trial cohort.
Click here to view more publications.
A substantial portion of people's risk of developing cancer is due to genetics. Much of the genetic variation underlying cancer risk affects regulatory elements (e.g., enhancers and promoters) that control gene expression. To understand how this variation in regulatory DNA confers risk of prostate cancer, we developed the cistrome-wide association study (CWAS). CWAS is a framework for identifying variants that impact traits and disease through their effects on chromatin state. CWAS associates the genetic determinants of cistromes (the genome-wide profiles of transcription factor binding sites or histone modifications) with phenotypes using summary statistics from genome-wide association studies (GWAS). We performed a CWAS of prostate cancer using a reference panel of 307 prostate cistromes from 165 individuals. CWAS identified hundreds of prostate cancer risk loci where regulatory element activity or transcription factor binding explain most of the inherited risk identified by GWAS. In addition, CWAS boosts discovery power in modestly sized GWAS, allowing us to identify novel genetic associations mediated through AR binding. This allowed us to discover a SNP associated with resistance to prostate cancer therapy from a clinical trial cohort.
Click here to view more publications.
HOME | RESEARCH | PEOPLE | PUBLICATIONS | SUPPORT | CONTACT
©2022 Baca Lab at Dana-Farber Cancer Institute
©2022 Baca Lab at Dana-Farber Cancer Institute
